Note
Go to the end to download the full example code or to run this example in your browser via Binder
FSLE experiment in med¶
Example to build Finite Size Lyapunov Exponents, parameter values must be adapted for your case.
Example use a method similar to AVISO flse
from matplotlib import pyplot as plt
from numba import njit
from numpy import arange, arctan2, empty, isnan, log, ma, meshgrid, ones, pi, zeros
from py_eddy_tracker import start_logger
from py_eddy_tracker.data import get_demo_path
from py_eddy_tracker.dataset.grid import GridCollection, RegularGridDataset
start_logger().setLevel("ERROR")
ADT in med¶
from_netcdf_cube() method is
made for data stores in time cube, you could use also
from_netcdf_list() method to
load data-cube from multiple file.
c = GridCollection.from_netcdf_cube(
get_demo_path("dt_med_allsat_phy_l4_2005T2.nc"),
"longitude",
"latitude",
"time",
# To create U/V variable
heigth="adt",
)
/home/docs/checkouts/readthedocs.org/user_builds/py-eddy-tracker/conda/latest/lib/python3.10/site-packages/numpy-1.24.3-py3.10-linux-x86_64.egg/numpy/ma/core.py:1020: RuntimeWarning: overflow encountered in multiply
result = self.f(da, db, *args, **kwargs)
Methods to compute FSLE¶
@njit(cache=True, fastmath=True)
def check_p(x, y, flse, theta, m_set, m, dt, dist_init=0.02, dist_max=0.6):
"""
Check if distance between eastern or northern particle to center particle is bigger than `dist_max`
"""
nb_p = x.shape[0] // 3
delta = dist_max**2
for i in range(nb_p):
i0 = i * 3
i_n = i0 + 1
i_e = i0 + 2
# If particle already set, we skip
if m[i0] or m[i_n] or m[i_e]:
continue
# Distance with north
dxn, dyn = x[i0] - x[i_n], y[i0] - y[i_n]
dn = dxn**2 + dyn**2
# Distance with east
dxe, dye = x[i0] - x[i_e], y[i0] - y[i_e]
de = dxe**2 + dye**2
if dn >= delta or de >= delta:
s1 = dn + de
at1 = 2 * (dxe * dxn + dye * dyn)
at2 = de - dn
s2 = ((dxn + dye) ** 2 + (dxe - dyn) ** 2) * (
(dxn - dye) ** 2 + (dxe + dyn) ** 2
)
flse[i] = 1 / (2 * dt) * log(1 / (2 * dist_init**2) * (s1 + s2**0.5))
theta[i] = arctan2(at1, at2 + s2) * 180 / pi
# To know where value are set
m_set[i] = False
# To stop particle advection
m[i0], m[i_n], m[i_e] = True, True, True
@njit(cache=True)
def build_triplet(x, y, step=0.02):
"""
Triplet building for each position we add east and north point with defined step
"""
nb_x = x.shape[0]
x_ = empty(nb_x * 3, dtype=x.dtype)
y_ = empty(nb_x * 3, dtype=y.dtype)
for i in range(nb_x):
i0 = i * 3
i_n, i_e = i0 + 1, i0 + 2
x__, y__ = x[i], y[i]
x_[i0], y_[i0] = x__, y__
x_[i_n], y_[i_n] = x__, y__ + step
x_[i_e], y_[i_e] = x__ + step, y__
return x_, y_
Settings¶
# Step in degrees for ouput
step_grid_out = 1 / 25.0
# Initial separation in degrees
dist_init = 1 / 50.0
# Final separation in degrees
dist_max = 1 / 5.0
# Time of start
t0 = 20268
# Number of time step by days
time_step_by_days = 5
# Maximal time of advection
# Here we limit because our data cube cover only 3 month
nb_days = 85
# Backward or forward
backward = True
Particles¶
FSLE¶
# Array to compute fsle
fsle = zeros(x0.shape[0], dtype="f4")
theta = zeros(x0.shape[0], dtype="f4")
mask = ones(x0.shape[0], dtype="f4")
x, y = build_triplet(x0, y0, dist_init)
used = zeros(x.shape[0], dtype="bool")
# advection generator
kw = dict(
t_init=t0, nb_step=1, backward=backward, mask_particule=used, u_name="u", v_name="v"
)
p = c.advect(x, y, time_step=86400 / time_step_by_days, **kw)
# We check at each step of advection if particle distance is over `dist_max`
for i in range(time_step_by_days * nb_days):
t, xt, yt = p.__next__()
dt = t / 86400.0 - t0
check_p(xt, yt, fsle, theta, mask, used, dt, dist_max=dist_max, dist_init=dist_init)
# Get index with original_position
i = ((x0 - x0_) / step_grid_out).astype("i4")
j = ((y0 - y0_) / step_grid_out).astype("i4")
fsle_ = empty(grid_shape, dtype="f4")
theta_ = empty(grid_shape, dtype="f4")
mask_ = ones(grid_shape, dtype="bool")
fsle_[i, j] = fsle
theta_[i, j] = theta
mask_[i, j] = mask
# Create a grid object
fsle_custom = RegularGridDataset.with_array(
coordinates=("lon", "lat"),
datas=dict(
fsle=ma.array(fsle_, mask=mask_),
theta=ma.array(theta_, mask=mask_),
lon=lon_p,
lat=lat_p,
),
centered=True,
)
/home/docs/checkouts/readthedocs.org/user_builds/py-eddy-tracker/conda/latest/lib/python3.10/site-packages/numpy-1.24.3-py3.10-linux-x86_64.egg/numpy/ma/core.py:4348: RuntimeWarning: invalid value encountered in multiply
self._data.__imul__(other_data)
/home/docs/checkouts/readthedocs.org/user_builds/py-eddy-tracker/conda/latest/lib/python3.10/site-packages/numpy-1.24.3-py3.10-linux-x86_64.egg/numpy/ma/core.py:4348: RuntimeWarning: invalid value encountered in multiply
self._data.__imul__(other_data)
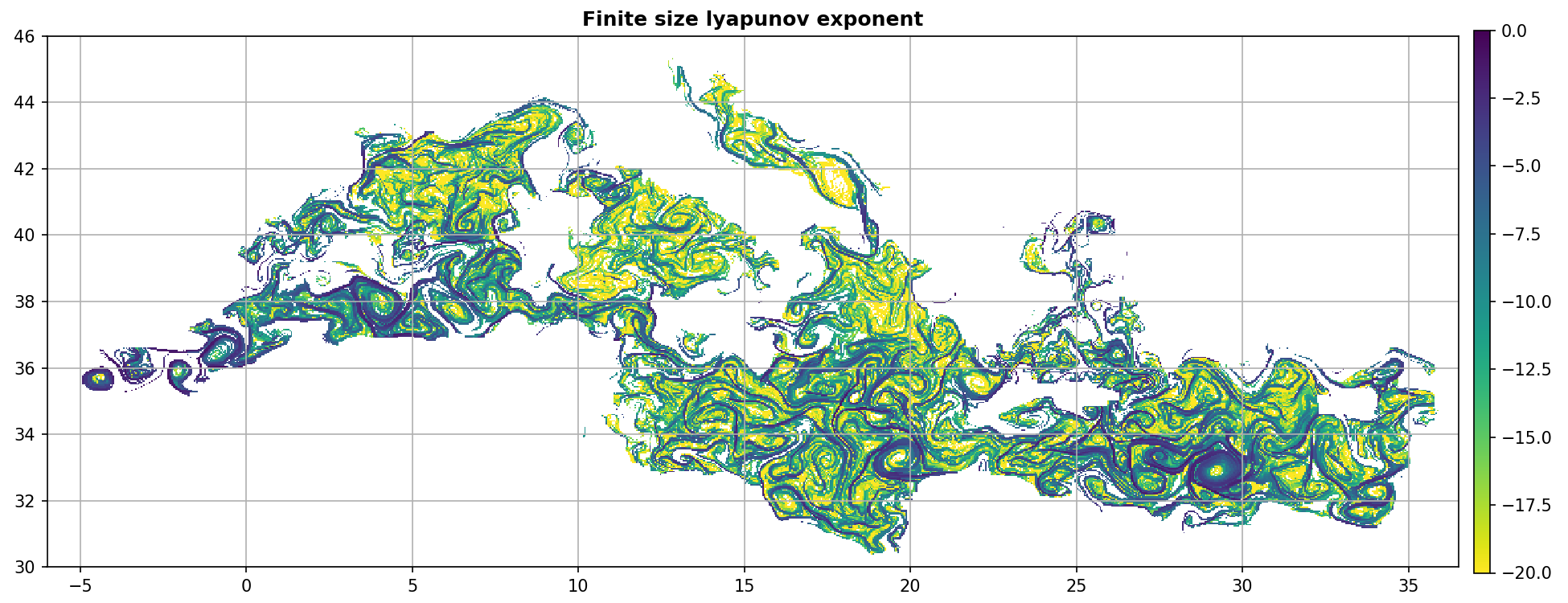
Display FSLE¶
fig = plt.figure(figsize=(13, 5), dpi=150)
ax = fig.add_axes([0.03, 0.03, 0.90, 0.94])
ax.set_xlim(-6, 36.5), ax.set_ylim(30, 46)
ax.set_aspect("equal")
ax.set_title("Finite size lyapunov exponent", weight="bold")
kw = dict(cmap="viridis_r", vmin=-20, vmax=0)
m = fsle_custom.display(ax, 1 / fsle_custom.grid("fsle"), **kw)
ax.grid()
_ = plt.colorbar(m, cax=fig.add_axes([0.94, 0.05, 0.01, 0.9]))
/home/docs/checkouts/readthedocs.org/user_builds/py-eddy-tracker/conda/latest/lib/python3.10/site-packages/numpy-1.24.3-py3.10-linux-x86_64.egg/numpy/ma/core.py:1163: RuntimeWarning: overflow encountered in divide
result = self.f(da, db, *args, **kwargs)
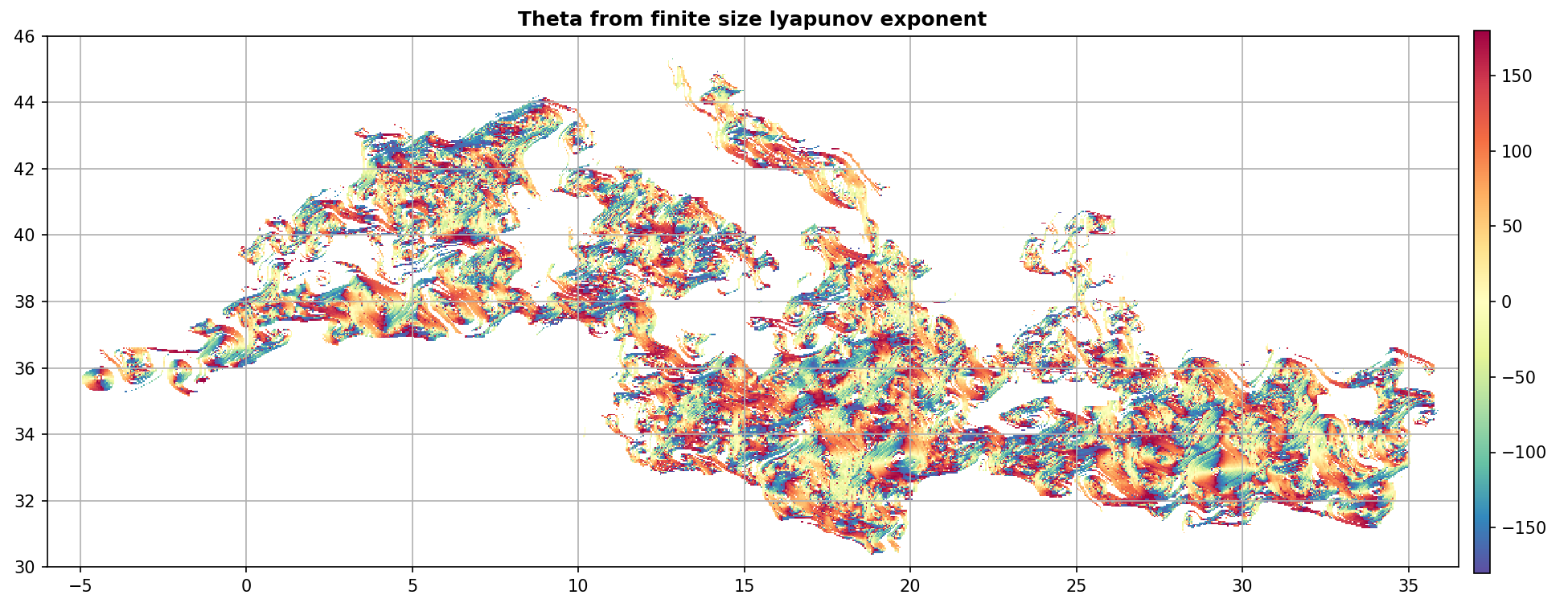
Display Theta¶
fig = plt.figure(figsize=(13, 5), dpi=150)
ax = fig.add_axes([0.03, 0.03, 0.90, 0.94])
ax.set_xlim(-6, 36.5), ax.set_ylim(30, 46)
ax.set_aspect("equal")
ax.set_title("Theta from finite size lyapunov exponent", weight="bold")
kw = dict(cmap="Spectral_r", vmin=-180, vmax=180)
m = fsle_custom.display(ax, fsle_custom.grid("theta"), **kw)
ax.grid()
_ = plt.colorbar(m, cax=fig.add_axes([0.94, 0.05, 0.01, 0.9]))
Total running time of the script: ( 0 minutes 11.798 seconds)