py_eddy_tracker.observations.observation.EddiesObservations¶
- class py_eddy_tracker.observations.observation.EddiesObservations(size=0, track_extra_variables=None, track_array_variables=0, array_variables=None, only_variables=None, raw_data=False)[source]¶
Bases:
objectClass to store eddy observations.
Methods
Add a new field.
Align the time indices of two datasets.
Merge.
Give major axis in km with a given latitude
- param str,array xname
variable to compute stats on
Return values evenly spaced with few numbers
Set contours as circles from radius and center data.
Check coherence between two datasets.
Return index of contour containing (x,y)
Copy with buffer for zarr.
Return the cost function between two obs.
How does it work on x bound ?
create particles only inside speed contour.
Plot the speed and effective (dashed) contour of the eddies
Use haversine distance for distance matrix between every self and other eddies.
Extract geographically with a bounding box.
Extract a subset of observations.
Produce description table of the fields available in this object
- param matplotlib.axes.Axes ax
matplotlib axe used to draw
Get first obs of each trajectory.
Return colors as a cyclic list
Get percentile of eddies in each bin
Count the eddies in each bin (use all pixels in each contour)
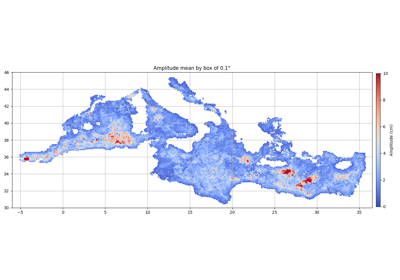
Return the mean of the eddies’ variable in each bin
Build histograms.
Return obs from self at the index.
Insert other obs in self at the given index.
True for each point inside the effective contour of an eddy
Interpolate a grid on a center or contour with mean, min or max method
Get flag of the eddy’s convexity
Yield observation group for each bin.
Get Last obs of each trajectory.
Load the netcdf or the zarr file.
Load data from netcdf.
Load data from zarr.
Return mask for all observations in one of polygons list
Return index and score computed on the effective contour.
Merge two datasets.
Compute an intersection between all filters after to evaluate each of them
Fill virtual obs (C).
Shift index with ref
Copy with fields listed remove
Scatter data.
Write something (TODO)
Time sub sampling
Track obs between self and other
Write a netcdf or zarr with eddy obs.
Attributes
Return dtype to build numpy array.
Return all the names of the variables.
Return period in days covered by the dataset
Return observations.
Give the time coverage.
- COLORS = ['sienna', 'red', 'darkorange', 'gold', 'palegreen', 'limegreen', 'forestgreen', 'mediumblue', 'dodgerblue', 'lightskyblue', 'violet', 'blueviolet', 'darkmagenta', 'darkgrey', 'dimgrey', 'steelblue']¶
- ELEMENTS = ['lon', 'lat', 'radius_s', 'radius_e', 'amplitude', 'speed_average', 'time', 'shape_error_e', 'shape_error_s', 'speed_area', 'effective_area', 'nb_contour_selected', 'num_point_e', 'num_point_s', 'height_max_speed_contour', 'height_external_contour', 'height_inner_contour']¶
- NB_COLORS = 16¶
- array_variables¶
- static basic_formula_ellipse_major_axis(lats, cmin=1.5, cmax=10.0, c0=1.5, lat1=13.5, lat2=5.0, degrees=False)[source]¶
Give major axis in km with a given latitude
- bins_stat(xname, bins=None, yname=None, method=None, mask=None)[source]¶
- Parameters
- Returns
x array and y array
- Return type
array,array
- circle_contour(only_virtual=False, factor=1)[source]¶
Set contours as circles from radius and center data.
- contains(x, y, intern=False)[source]¶
Return index of contour containing (x,y)
- Parameters
x (array) – longitude
y (array) – latitude
intern (bool) – If true use speed contour instead of effective contour
- Returns
indexs, -1 if no index
- Return type
array[int32]
- static copy_data_to_zarr(handler_zarr, handler_eddies, sl_obs, buffer_size, factor, raw_data, scale_factor, add_offset)[source]¶
Copy with buffer for zarr.
Zarr need to get real value, and size could be huge, so we use a buffer to manage memory :param zarr_dataset handler_zarr: :param array handler_eddies: :param slice zarr_dataset sl_obs: :param int buffer_size: :param float factor: :param bool raw_data: :param None,float scale_factor: :param None,float add_offset:
- static cost_function(records_in, records_out, distance)[source]¶
Return the cost function between two obs.
\[cost = \sqrt{({Amp_{_{in}} - Amp_{_{out}} \over Amp_{_{in}}}) ^2 + ({Rspeed_{_{in}} - Rspeed_{_{out}} \over Rspeed_{_{in}}}) ^2 + ({distance \over 125}) ^2 }\]- Parameters
records_in – starting observations
records_out – observations to associate
distance – computed between in and out
- classmethod cost_function_common_area(xy_in, xy_out, distance, intern=False)[source]¶
How does it work on x bound ?
- Parameters
xy_in –
xy_out –
distance –
intern (bool) –
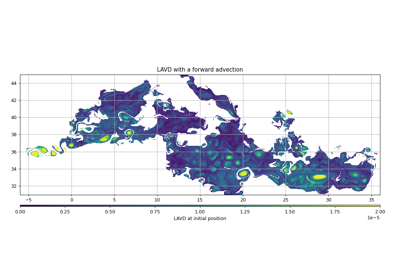
- create_particles(step, intern=True)[source]¶
create particles only inside speed contour. Avoid creating too large numpy arrays, only to me masked
- create_variable(handler_nc, kwargs_variable, attr_variable, data, scale_factor=None, add_offset=None, **kwargs)[source]¶
- create_variable_zarr(handler_zarr, kwargs_variable, attr_variable, data, scale_factor=None, add_offset=None, filters=None, compressor=None, chunck_size=2500000)[source]¶
- display(ax, ref=None, extern_only=False, intern_only=False, **kwargs)[source]¶
Plot the speed and effective (dashed) contour of the eddies
- Parameters
ax (matplotlib.axes.Axes) – matplotlib axe used to draw
extern_only (bool) – if True, draw only the effective contour
intern_only (bool) – if True, draw only the speed contour
kwargs (dict) – look at
matplotlib.axes.Axes.plot()
- distance(other)[source]¶
Use haversine distance for distance matrix between every self and other eddies.
- property dtype¶
Return dtype to build numpy array.
- property elements¶
Return all the names of the variables.
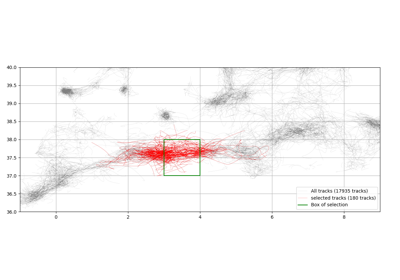
- extract_with_area(area, **kwargs)[source]¶
Extract geographically with a bounding box.
- Parameters
area (dict) – 4 coordinates in a dictionary to specify bounding box (lower left corner and upper right corner)
kwargs (dict) – look at
extract_with_mask()
- Returns
Return all eddy trajetories in bounds
- Return type
area = dict(llcrnrlon=x0, llcrnrlat=y0, urcrnrlon=x1, urcrnrlat=y1)
- extract_with_mask(mask)[source]¶
Extract a subset of observations.
- Parameters
mask (array(bool)) – mask to select observations
- Returns
same object with selected observations
- Return type
self
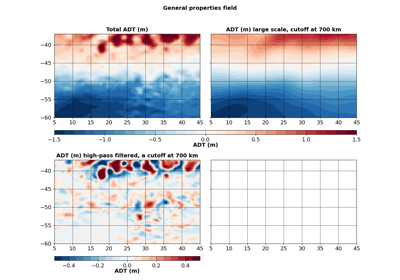
- filled(ax, varname=None, ref=None, intern=False, cmap='magma_r', lut=10, vmin=None, vmax=None, factor=1, **kwargs)[source]¶
- Parameters
ax (matplotlib.axes.Axes) – matplotlib axe used to draw
varname (str,array,None) – variable used to fill the contours, or an array of same size than obs
ref (float,None) – if defined, all coordinates are wrapped with ref as western boundary
intern (bool) – if True draw speed contours instead of effective contours
cmap (str) – matplotlib colormap name
factor (float) – multiply value by
- Returns
Collection drawed
- Return type
- property global_attr¶
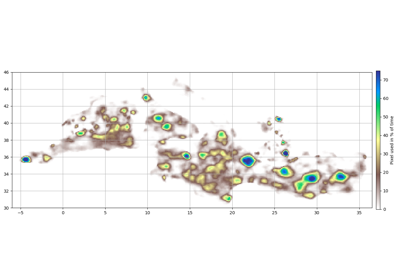
- grid_box_stat(bins, varname, method=50, data=None, filter=slice(None, None, None))[source]¶
Get percentile of eddies in each bin
- Parameters
- Returns
return grid of method
- Return type
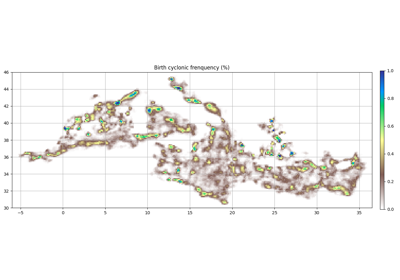
- grid_count(bins, intern=False, center=False, filter=slice(None, None, None))[source]¶
Count the eddies in each bin (use all pixels in each contour)
- Parameters
- Returns
return the grid of counts
- Return type
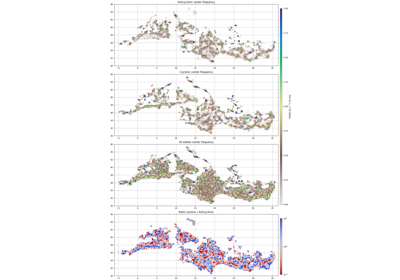
- grid_stat(bins, varname, data=None)[source]¶
Return the mean of the eddies’ variable in each bin
- Parameters
bins ((numpy.array,numpy.array)) – bins (grid) to compute the mean on
varname (str) – name of variable to compute the mean on and output grid_name
data (array) – Array used to compute stat if defined
- Returns
return the gridde mean variable
- Return type
- interp_grid(grid_object, varname, method='center', dtype=None, intern=None)[source]¶
Interpolate a grid on a center or contour with mean, min or max method
- Parameters
grid_object (py_eddy_tracker.dataset.grid.RegularGridDataset) – Handler of grid to interp
varname (str) – Name of variable to use
method (str) – ‘center’, ‘mean’, ‘max’, ‘min’, ‘nearest’
dtype (str) – if None we use var dtype
intern (bool) – Use extern or intern contour
- iter_on(xname, bins=None)[source]¶
Yield observation group for each bin.
- Parameters
xname (str,array) –
bins (array) – bounds of each bin ,
- Returns
index or mask, bound low, bound up
- classmethod load_file(filename, **kwargs)[source]¶
Load the netcdf or the zarr file.
Load only latitude and longitude on the first 300 obs :
kwargs_latlon_300 = dict( include_vars=["longitude", "latitude",], indexs=dict(obs=slice(0, 300)), ) small_dataset = TrackEddiesObservations.load_file( filename, **kwargs_latlon_300 )
For **kwargs look at
load_from_zarr()orload_from_netcdf()
- classmethod load_from_netcdf(filename, raw_data=False, remove_vars=None, include_vars=None, indexs=None, **class_kwargs)[source]¶
Load data from netcdf.
- Parameters
filename (str,ExFileObject) – path or handler to load data
raw_data (bool) – If true load data without apply scale_factor and add_offset
remove_vars (None,list(str)) – List of variable name which will be not loaded
include_vars (None,list(str)) – If defined only this variable will be loaded
class_kwargs – argument to set up observations class
- Returns
Obsevations selected
- Return type
class
- classmethod load_from_zarr(filename, raw_data=False, remove_vars=None, include_vars=None, indexs=None, buffer_size=5000000, **class_kwargs)[source]¶
Load data from zarr.
- Parameters
filename (str,store) – path or store to load data
raw_data (bool) – If true load data without scale_factor and add_offset
remove_vars (None,list(str)) – List of variable name that will be not loaded
include_vars (None,list(str)) – If defined only this variable will be loaded
buffer_size (int) – Size of buffer used to load zarr data
class_kwargs – argument to set up observations class
- Returns
Obsevations selected
- Return type
class
- mask_from_polygons(polygons)[source]¶
Return mask for all observations in one of polygons list
- Parameters
polygons (list((array,array))) – list of x/y array which be used to identify observations
- match(other, i_self=None, i_other=None, method='overlap', intern=False, cmin=0, **kwargs)[source]¶
Return index and score computed on the effective contour.
- Parameters
other (EddiesObservations) – Observations to compare
i_self (array[bool,int],None) – Index or mask to subset observations, it could avoid to build a specific dataset.
i_other (array[bool,int],None) – Index or mask to subset observations, it could avoid to build a specific dataset.
method (str) –
“overlap”: the score is computed with contours;
”circle”: circles are computed and used for score (TODO)
intern (bool) – if True, speed contour is used (default = effective contour)
cmin (float) – 0 < cmin < 1, return only couples with score >= cmin
kwargs (dict) – look at
vertice_overlap()
- Returns
return the indices of the eddies in self coupled with eddies in other and their associated score
- Return type
- merge_filters(*filters)[source]¶
Compute an intersection between all filters after to evaluate each of them
- property nb_days¶
Return period in days covered by the dataset
- Returns
Number of days
- Return type
- property obs¶
Return observations.
- observations¶
- only_variables¶
- property period¶
Give the time coverage. If collection is empty, return nan,nan
- period_¶
- propagate(previous_obs, current_obs, obs_to_extend, dead_track, nb_next, model)[source]¶
Fill virtual obs (C).
- Parameters
previous_obs – previous obs from current (A)
current_obs – previous obs from virtual (B)
obs_to_extend –
dead_track –
nb_next –
model –
- Returns
New position C = B + AB
- raw_data¶
- scatter(ax, name=None, ref=None, factor=1, **kwargs)[source]¶
Scatter data.
- Parameters
ax (matplotlib.axes.Axes) – matplotlib axe used to draw
name (str,array,None) – variable used to fill the contour, if None all elements have the same color
ref (float,None) – if defined, all coordinates are wrapped with ref as western boundary
factor (float) – multiply value by
kwargs (dict) – look at
matplotlib.axes.Axes.scatter()
- Returns
scatter mappable
- property shape¶
- property sign_legend¶
- sign_type¶
- track_array_variables¶
- track_extra_variables¶
- property tracks¶